 These cases also could be fully displayed and visualized in the heat map. Moreover, the STE amplitude in 40% (24 out of 60) of cases reached the threshold specified in the STEMI guideline. According to cluster analysis in the heat map, STE leads were clustered into two categories, comprising of the right precordial leads (V 1, V 2, V 3) and others (V 4, V 5, V 6, I, II, III, aVF, aVL, aVR). 
The ST-segment shifts of each lead of each collected ECG could be conveniently visualized in the heat map. STE leads were mainly in the V 1, V 2 and V 3 leads. In total, 60 cases of electrocardiographic LVH with STE were screened and analyzed. Cluster analysis was carried out based on the heat map and the results were drawn as tree maps (pedigree maps) in the heat map. HemI 1.0 software was used to draw heat maps to display the STE of each lead of each collected ECG. We sequentially collected the electrocardiograms of inpatients in the First Affiliated Hospital of Shantou University Medical College from July 2015 to December 2015 in order to screen cases of LVH with STE. The aim of this study was to analyze and show data for electrocardiographic left ventricular hypertrophy (LVH) with ST-segment elevation (STE) by a heat map in order to explore the feasibility and clinical value of heat mapping for ECG data visualization. Few papers display ECG data by visual means. Heatmap was plotted by, a free online platform for data analysis and visualization.Most electrocardiogram (ECG) studies still take advantage of traditional statistical functions, and the results are mostly presented in tables, histograms, and curves. You can cite the original package, or using the following format: Script need strigent input format, please read the instructions and examples carefully.Ĩ00+ papers cited ( Google Scholar). Scale indicates different thresholds of the p-value, and the size of the dot indicates the number of genes corresponding to each pathway.Ģ, Open with excel, and change into the same format as the exampleģ, Copy and paste data into the input frame The horizontal axis represents the gene ratio, while the vertical axis represents the enriched pathway name. Honghua extract mediated potent inhibition of COVID-19 host cell pathwaysĭot plot of the KEGG pathway enrichment analysis. Optional for fifth columns, plot by class with facet, for example BP, CC, MF. The fourth colunn is gene count (size of bubble). The third column is p (or FDR), represents colors 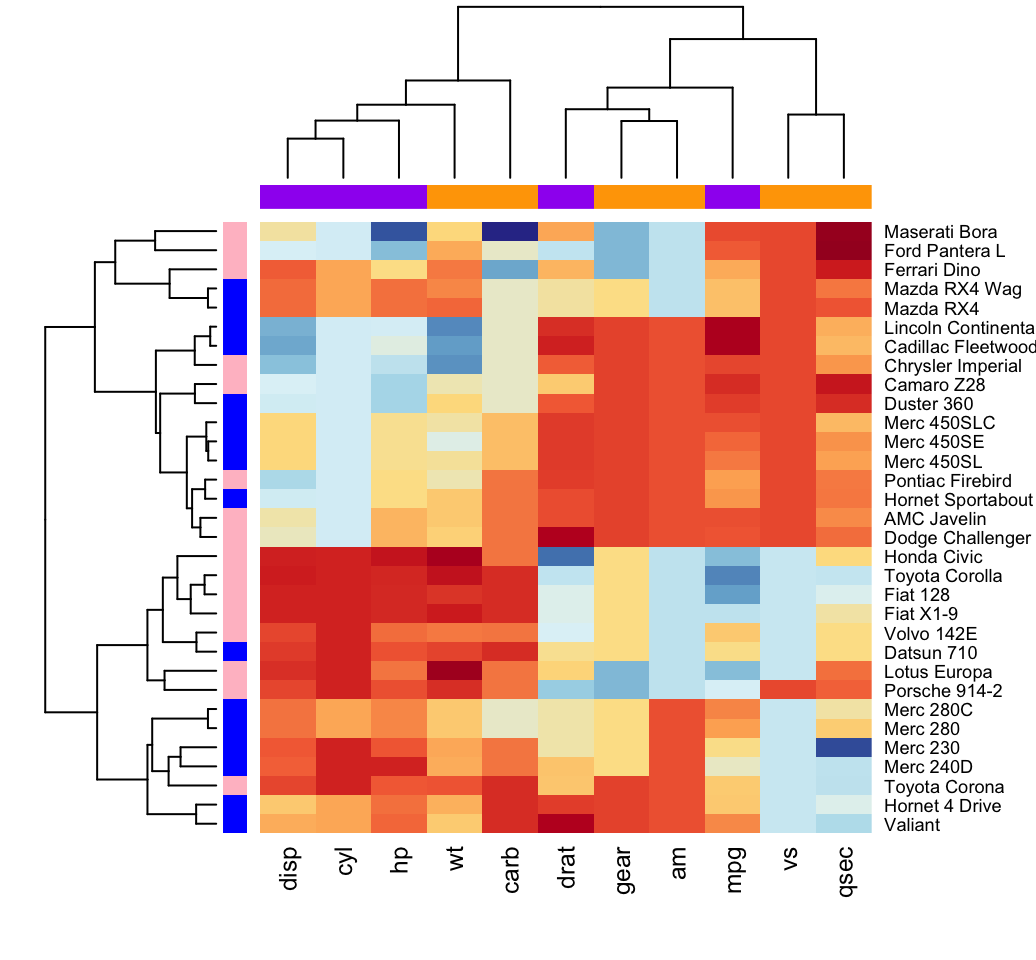
The second column is enrichment (display in X, if no, replace by others, such as gene ratio, -log10(p), count Input data contain 4 columns: the first column is GO or KEGG Pathway term (display in Y axis) Please use GO, Pathway Enrichment Analysis or metascape to get the enrichment results, and then plot. 
GO, Pathway enrichment bubble plot Introductionīubble plot is generally used in GO, KEGG pathway enrichment analysis, in which p values are represented by colors, gene counts are represented by bubble size. Larger -log10(pvalue) / smaller pvalue color:Ĭolumn 2, big -> small column 2, small -> bigĬolumn 3, big -> small column 3, small -> bigĬolumn 4, big -> small column 4, small -> big Smaller -log10(pvalue) / larger pvalue color: 
If choose pvlaue in colors options, labbled as pvalueĬolors (take pvalue as an example, can also be FDR, qvalue. If choose -log10 transform in colors options, labled as -log10(pvalue) Note: input data format must match the example on the right, tab-seperatedġ, data from excel, copy and paste data into the input frameĢ, data from txt, must tab-seperated, copy and paste data into the input frameģ, use point as decimal separator, not comma. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
